Products
Innovative Yeast-Based And Molecular Biology Kits To Accelerate Your Life Sciences Research.
PRODUCTS


Saccharomyces Cerevisiae Surface Display Vector Kit

Price:
The Saccharomyces Cerevisiae Surface Display Vector Kit offers an efficient solution for protein expression and functional screening in yeast. Includes EBY100 strain, pYD1 vector, control strains, and primers for protein research. Buy now!
INSTRUCTION & COA:
- Product Description
-
Product Overview
The Saccharomyces cerevisiae surface display system is an efficient method for displaying and analyzing exogenous proteins on the surface of yeast cells. This versatile technique is widely used for protein expression, functional screening, and interaction studies in biotechnology and pharmaceutical research. Our Saccharomyces cerevisiae Surface Display Vector Kit provides everything necessary to introduce a target exogenous protein gene into yeast cells for surface expression.
This kit utilizes the Agglutinin yeast display system, one of the most commonly employed systems for yeast surface display. The agglutinin complex, composed of Aga1p and Aga2p subunits, facilitates efficient display of recombinant proteins on the surface of Saccharomyces cerevisiae. This kit includes the EBY100 yeast strain, the pYD1 vector, positive and empty control strains, as well as the necessary primers for amplification and sequencing (PYD1-F and PYD1-R).
CAT: RY8019
Price and Instruction: Available upon inquiry
Product Components
Component
Name
Concentration
Specification
Quantity
Plasmids
pYD1
100 ng/µL
20 µL
1 tube
Strain
EBY100
Inoculated Plate
Diameter: 90mm
1 plate
EBY100-pYD1-Empty control strain
Inoculated Plate
Diameter: 90mm
1 plate
EBY100-pYD1-Positive control strain
Inoculated Plate
Diameter: 90mm
1 plate
Primers
pYD1-F
Lyophilized powder
2 OD
1 tube
pYD1-R
Lyophilized powder
2 OD
1 tube
Storage and Shipping Conditions
Plasmids: Store at -20°C, shipped at room temperature.
Strains: Store at room temperature, shipped at room temperature.
Primers: Store at -20°C, shipped at room temperature.
Experimental Principle
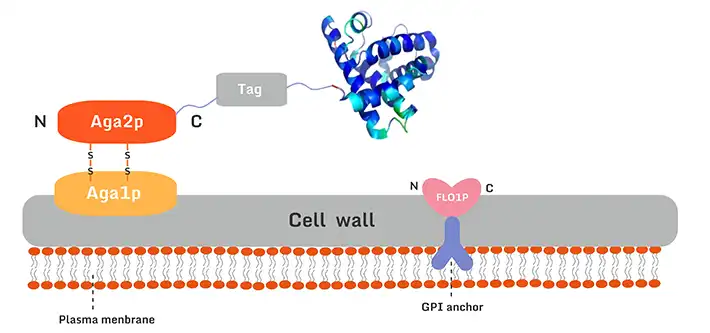
The Saccharomyces cerevisiae surface display involves the fusion of a target exogenous protein gene with specific vector gene sequences, which are then introduced into yeast cells. Using the yeast's natural protein transport mechanism, the target protein is transported to and anchored on the yeast cell surface.
The Agglutinin yeast display system includes two essential components: the core subunit Aga1p and the binding subunit Aga2p, enabling stable and efficient display of recombinant proteins. This system is ideal for applications such as protein-protein interaction studies, protein functional screening, and ligand binding assays.
Experimental Guidelines
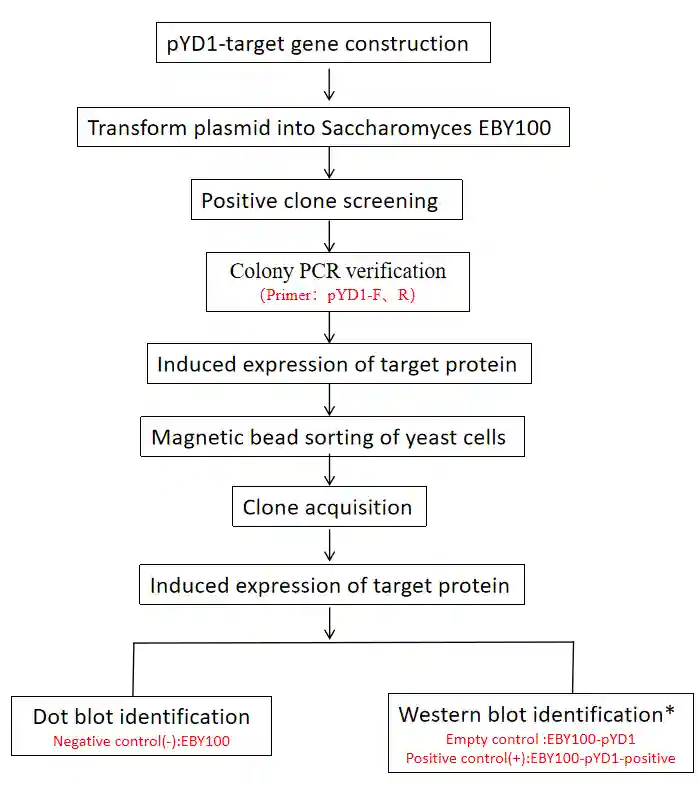
Note: Empty control strain EBY100-pYD1: 40 kDa; Positive control strain EBY100-pYD1-Positive: 95 kDa.
Precautions
1. To ensure strain viability, streak the strain onto a YPDA agar plate before liquid culturing.
2. Store strains at -80°C with 50% glycerol to prevent repeated freeze-thaw cycles.
3. This product is for research use only and not for clinical, diagnostic, or therapeutic applications.
4. Maintain sterile conditions to avoid contamination.
FAQ
Q: What is Saccharomyces cerevisiae surface display used for?
A: Saccharomyces cerevisiae surface display is widely used for studying protein interactions, functional screening, and protein expression in yeast. It is an essential technique in biotechnology, pharmaceutical research, and protein engineering.Q: What are the components of the Saccharomyces cerevisiae Surface Display Vector Kit?
A: The kit includes the EBY100 yeast strain, pYD1 vector, positive and empty control strains, and primers for amplification and sequencing (PYD1-F and PYD1-R).Ready to Advance Your Research?
Contact us today for a detailed consultation and product quotation. Let us help you unlock new insights your research. When inquiring about this product, please reference Product Code: RY8019.
Statement: All products are for research purposes only. Not for use in diagnostic or therapeutic applications.
关键词:- Advanced Saccharomyces cerevisiae Surface Display

Contact Us

Tel: +86 025-85205672

Email: info@pronetbio.com

Address: Building 3C, Nanjing Xianlin Zhigu,
Qixia District, Nanjing, Jiangsu, China, 210033

Need more info?
Let's connect!


